Many people have an illness that is affected by changes in one or many of their genes. Researchers are studying these genetic changes, or “genomic variants,” to better understand the role that genetics play in health and disease.
What is a genomic variant?
All humans have near-identical DNA sequences. The slight differences that exist between everyone’s DNA sequences is what makes each of us unique. These differences, called genomic variants, occur within our DNA.
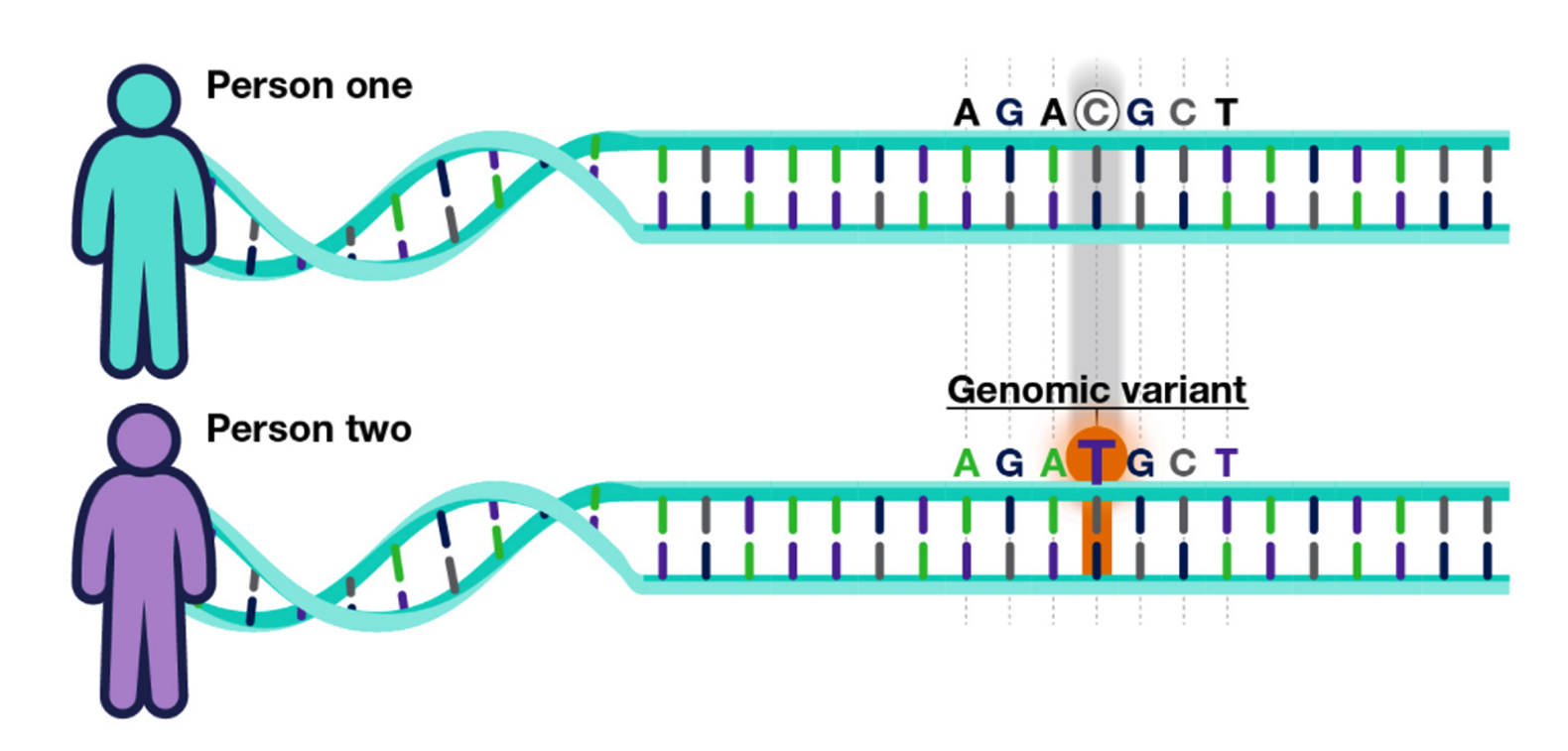
DNA is read like a code. This code is made up of 4 types of chemical building blocks: adenine, thymine, cytosine, and guanine, abbreviated with the letters A, T, C and G. A genomic variant is when a person’s DNA code, in a certain location, differs from other peoples’ DNA code in that same location.
In this example, Person one has a “C” whereas Person two, has a “T” in the same location.
 There are roughly 4 to 5 million genomic variants in each person’s genome. These variants (or differences) may or may not be unique to a person. Some variants may increase a person’s risk of developing diseases, others may reduce risk, and some have no effect at all.
There are roughly 4 to 5 million genomic variants in each person’s genome. These variants (or differences) may or may not be unique to a person. Some variants may increase a person’s risk of developing diseases, others may reduce risk, and some have no effect at all.
Using Genomic Variants to Create Polygenic Risk Scores
To identify genomic variants related to diseases, researchers compare genomes of people with and without those diseases and use statistics to estimate how these variants together may affect individuals’ risk for certain diseases. These calculations create polygenic risk scores.
Polygenic Risk Scores and KP Research Bank Data
Two recently approved KP Research Bank projects highlighted in this newsletter show how researchers are using Research Bank data to develop polygenic risk scores that may serve as an important new tool to guide health care decisions for people from all backgrounds.
For more information on polygenic risk scores, watch this video from the National Human Genome Research Institute.
(Source: “Polygenic Risk Scores,” National Human Genome Research Institute, genome.gov/Health/Genomics-and-Medicine/Polygenic-risk-scores, August 11, 2020.)


